No products
Product successfully added to your shopping cart
There are 0 items in your cart. There is 1 item in your cart.
Molecular Probes
- Protein Control Ligand
- Pathway Inhibitors
- Enzyme Inhibitors
- Kinase Inhibitors
- Protease
- Synthase
- p18
- p38
- p53
- p70
- p90
- Peptidase
- Carboxyl and Decarboxylases
- Ceramide Turnover Enzymes
- Chromatin Modifying Enzymes
- Cyclic Nucleotide Turnover Enzymes
- Glycerophospholipid Turnover Enzymes
- Hydroxylases
- Ubiquitin-Activating Enzyme
- Adenosine Deaminase
- Clathrin
- Nuclease
- p68
- ACE
- COX
- DHFR
- Neprilysin
- NF-κB
- RAF
- RAS
- Reductase
- ROR
- Topoisomerase
- Transferase
- Protein Inhibitors
- Transporter Inhibitors
- Cell Inhibition
- Synthase
- Receptor Tyrosine Phosphatases (RTP)
- AChE
- Peptidase
- Autophagy
- Toll-Like Receptor (TLR)
- Enzyme Inhibitors
- Function Modulators
- Activators
- G Protein-Coupled Receptor Ligands
- 5HT Receptors
- Adrenoceptor
- Angiotensin Receptor
- Cannabinoid Receptors
- CCK Receptors
- DA Receptors
- EAA Receptors
- Ghrelin Receptors
- GABA Receptors
- Histamine Receptors
- Leukotriene Receptors
- Metabotropic Glutamate Receptors
- Motilin Receptors
- Muscarinic Receptor
- Neuropeptide Receptors
- Opioid Receptors
- Orexin Receptors
- Orphan Receptors
- Prostanoid Receptors
- Proteinase-Activated Receptors
- Purinergic Receptors
- Ryanodine receptor
- Sigma Receptors
- Thrombin Receptor
- Vaniloid Receptor
- VIP and PACAP Receptors
- Neurotensin Receptors
- Urotensin Receptor
- Imidazoline receptor
- SMO Receptors
- Apelin Receptor
- β-arrestin/β2-adaptin
- KDM4
- Glucocorticoid Receptor
- Laminin Receptor
- AHR
- Amylin Receptor
- Bombesin Receptor
- Bradykinin Receptor
- CFTR
- CGRP Receptor
- CRFR
- Endothelin Receptor
- Ephrin Receptor
- Farnesoid X receptor (FXR)
- Glucagon Receptor
- Nuclear Receptor Ligands
- GDNF Receptors
- TNF Receptors
- Transcription Factors
- Chemokines
- Cytokine Receptors
- Biomarkers and Buffer Solutions
- Molecular Probes
- Stem Cell Research
- Alzheimer's Disease
- Apoptosis
- Cancer Research
- Epigenetics
- Metabolites
- PET/SPECT Imaging Precursors
- Customized Screening Library
- Ultra Pure Pharmacological Standard
- Tissue Microarray (TMA)
- Proteins and Antibodies
- Primary Cells
- ELISA KIT
- Natural Products
- Lab Equipments
- Humanized Mice for PDX Platform
- Rare Chemicals
- Custom Synthesis
- Antibacterial
- Antifungal
- Antioxidant
- Antiviral
- Molecular Glues
- PROTAC Linker
- SARS-CoV
 View larger
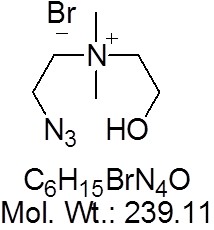
View larger 1-Azidoethylcholine Bromide
AOB7337
CAS: 2059973-54-3 (aminium: 1239916-55-2)
Chemical Name: AzCho; AECho; Azidoethyl-Cho; N-(2-Azidoethyl)-2-hydroxy-N,N-dimethyl-ethanaminium bromide
7932 Items
Molarity Calculation Cart®
HOW TO ORDER
Quantity Discount Table - Order More To Get More Price Discount
| Quantity | mg | Unit Price ($/mg or $/Unit) | Final Price |
|---|---|---|---|
| 1 | 5 | $31.45 | Total: $157.25 |
| 1 | 10 | $26.64 | Total: $266.40 |
| 1 | 25 | $22.57 | Total: $564.25 |
| 1 | 50 | $19.24 | Total: $962.00 |
| 1 | 100 | $16.65 | Total: $1,665.00 |
Data sheet
| Molecular Formula | C6H15BrN4O |
| Molecular Weight | 239.11 |
| CAS Numbers | 2059973-54-3 (aminium: 1239916-55-2) |
| Storage Condition | 0°C (short term), -20°C (long term), desiccated |
| Solubility | DMSO and Water |
| Purity | 98% by HPLC |
| Synonym | AzCho; AECho; Azidoethyl-Cho |
| IUPAC/Chemical Name | N-(2-Azidoethyl)-2-hydroxy-N,N-dimethyl-ethanaminium bromide |
| SMILES Code | [N-]=[N+]=NCC[N+](C)(CCO)C.[Br-] |
| References | 1) John T Ngo et al., Click-EM for imaging metabolically tagged nonprotein biomolecules; Nature Chemical Biology volume 12, pages459–465 (2016) |
More info
Choline phospholipids labeling agent, visualizing the subcellular localization of choline phospholipids.
What is it?
It is a choline analogue: the molecule mimics the head-group of choline (which is incorporated into phospholipids like phosphatidylcholine). ResearchGate+3salic.med.harvard.edu
It has an azido (-N₃) group attached (hence “azidoethyl-choline”) which renders it “clickable” (able to participate in bioorthogonal chemistry). salic.med.harvard.edu+2
One salt variant is the bromide form. aobious.com+1
What is the functionality / how is it used?
Here are the main functionalities and applications:
Metabolic incorporation into choline-containing phospholipids
Cells naturally take up choline, activate it (via phosphorylation, CDP-choline path) and incorporate it into phosphatidylcholine (PC) and related lipids. PMC+1
The azidocholine analogue is taken up similarly, incorporated into choline-phospholipids (for example into PC) such that the resulting lipid bears the azide tag. For example: AECho → “azidoethyl-phosphatidylcholine” in cells. salic.med.harvard.edu+1
Because the analogue is tolerated (i.e., doesn’t majorly perturb normal lipid pools) and integrates into the membranes, it gives a way to trace or image choline-lipid metabolism/dynamics. salic.med.harvard.edu+1
Bioorthogonal “click chemistry” labelling
Once the azide-tagged lipid is in place in a cell or organism, one can apply a complementary reagent (for example alkyne-fluorophore, dibenzocyclooctyne (DBCO)-fluorophore, etc) to “click” onto the azide and thus tag the lipid with a fluorophore or probe. salic.med.harvard.edu+1
This enables visualization of the labeled lipids: imaging their location (cell membrane surface, organelles), their trafficking, dynamics. For example the 2015 study by Jao et al. showed localization to mitochondria, internal membranes etc. salic.med.harvard.edu
It also opens possibility for two-colour labelling: e.g., combining azidocholine (azide) with propargyl-choline (alkyne) and then using orthogonal click reactions to differentiate two pools. salic.med.harvard.edu+1
Monitoring phospholipid synthesis / cellular membrane studies
Because it labels choline-phospholipids (which are abundant and functionally critical in membranes), this reagent is used in studies of membrane biology, lipid metabolism, membrane trafficking, organelle biology. For example: monitoring choline-containing phospholipid synthesis in cell culture and even in whole organisms.
More recently, it has been used in bacterial studies: e.g., metabolic labeling in bacteria (gram negative) to assess outer-membrane integrity via phospholipid externalization. ScienceDirect+1
Key properties
Molecular formula (for the cation): C₆H₁₅N₄O (for the iodo salt, but similar for bromide)
Molecular weight: ~159.21 g/mol for the cation in some form.
Solubility: Water / PBS buffer.
Storage: Typically –20 °C, avoid freeze/thaw.
Purity: ≥ 95% (H NMR confirms) for reagent quality.
Advantages & strengths
Allows specific labelling of choline-headgroup lipids with minimal perturbation of the natural lipidome (in the right conditions) — the Jao et al. paper found that AECho was incorporated without grossly changing non-choline phospholipids. salic.med.harvard.edu
Enables high‐resolution imaging by combining metabolic labeling + click chemistry + fluorescence. Gives insight into localization, trafficking of choline lipids in live or fixed cells.
Because it uses the endogenous metabolism of choline, it can serve as a reporter of biosynthetic activity of choline lipids, or changes in that metabolism.
The azide functionality is bioorthogonal (i.e., minimally interfering with native biology) which means the system can be relatively benign.
Limitations / considerations
Since it is a modified analogue of choline, if used at too high concentrations it could perturb the lipid metabolism or cell physiology. One must optimize labelling concentration and time. (Indeed the literature assessed perturbation of lipid pools). salic.med.harvard.edu
The click reaction (e.g., DBCO or SPAAC) requires reagents and perhaps fixation or specific conditions; live‐cell compatibility may have constraints (e.g., cytotoxicity of reagents, background labelling).
Because it competes with endogenous choline, if choline levels are high or the analogue uptake is low, the incorporation may be less efficient. For example, Jao et al. noted ~20% replacement of total choline‐PC at high analogue conc and 24 h. salic.med.harvard.edu
It labels choline-containing phospholipids, so it’s specific to that class; it does not label all lipids. One must interpret accordingly.
It is a research reagent (RUO – research use only) and not intended for therapeutic or clinical diagnostic use.
Example use‐case
From Jao et al. (2015): They incubated mammalian cells with AECho, then isolated lipids, and showed by mass spectrometry that the analogue was incorporated into phosphatidylcholine classes without disturbing other phospholipids. They then used a fluorescent DBCO–Alexa488 reagent to click onto the azide and imaged sub‐cellular localisation: cell surface membranes, organelles, mitochondria. salic.med.harvard.edu
Another study in bacteria used AECho metabolic labelling as a probe of outer‐membrane (OM) integrity: in mutants with compromised OM, the labelled phospholipids became accessible to the fluorescent click reagent, enabling detection of membrane defects. ScienceDirect
Practical tips for lab use
Optimize concentration and incubation time: too low → weak labelling; too high → possible perturbation.
Ensure the click reagent is compatible (e.g., strain-promoted alkyne reagent if doing live‐cell, to avoid copper toxicity).
Consider controls: unlabeled cells, choline competition, or blocking uptake to assess specificity.
After labelling, one can analyse by fluorescence microscopy, flow cytometry, or lipidomic (mass spec) analysis.
For imaging, ensure the fluorophore choice and imaging conditions match the expected membrane localisation (e.g., exclude nuclear signal, verify membrane vs cytosol).
Because the reagent is “head‐group” analogue of choline, confirm that downstream functional perturbation is minimal in your cell system (especially if studying metabolic/structural lipid biology).